G4 FUNNIES
G4 FUNNIES is a dedicated webserver engineered for the classification of proteins as G-Quadruplex-RNA binders.
The genesis of this algorithm is rooted in the training process, where it learned to differentiate between nuclear proteins that associate with folded G4 and those that bind to unfolded G4. The distinguishing environmental conditions, rich in Potassium (K+) and Lithium (Li+), facilitated the distinct structuring and unstructuring of G4, respectively.
The acronym G4 FUNNIES encapsulates the core components and objectives of the tool:
F: Folded - Representing the state of G4 RNA in a Potassium-rich environment.
UN: UNfolded - Denoting the state of G4 RNA in a Lithium-rich setting.
N: Nuclear - Signifying the nuclear proteins the tool was trained on
I: Interactome - Highlighting the focus on interactions between proteins and G4 RNA.
E: Explorer - Indicating the tool's exploratory nature in identifying and classifying these interactions.
S System - Stressing the organized and systematic method of classification.
By analyzing the binding of proteins in these contrasting environments, G4 FUNNIES not only identifies but also classifies G-Quadruplex-RNA binders with precision.
Introduction to the G4-Binding Protein Predictor
The tool uses the models created by the cleverMachine approach on two reference datasets. These datasets, named positive and negative sets, contain Li+ and K+ proteins respectively and were analysed by cleverMachine to detect the optimal combination of physicochemical scales that better discriminates them. All the predictive information of each scale is then combined together to form a complete model which is able to assign a new dataset to either be closer to positive (Li+) or negative (K+) class of the model. More detailed information on the derived model can be found here.
Internal workings
Input data
The G4-Binding Protein Predictor takes a FASTA dataset as an input in either text or file format.
Pre-filtering Customization

The prefilter for proteins with RNA-binding ability is built using our algorithm catRAPID signature. The method identifies proteins able to interact with RNAs. More details are available at the documentation page. The prefilter for protein with G4-binding ability utilizes an algorithm based on the protein set that consistently ranks lowest in our G4 mass spectrometry data.
In Figure a) we show that when eCLIP-identified RBP target occurrences of G4 RNA are processed using the G4 RNA propensity score of pqsfinder,
a clear distinction emerges between G4 binders and non-binders according to the cleverMACHINE classification (total of 150 RBPs).
This division is notably pronounced with elevated scores from the G4 RNA prediction tool, underscoring the ability of our cleverMACHINE in segregating the two groups.
Figure b) further emphasizes this distinction, revealing that as the confidence score of the cleverMACHINE escalates, a corresponding increase in the detection of G4 RNA is observed across the two protein categories
(those that bind to G4 RNA versus those that do not).
Such a pattern attests to our model's capability to precisely pinpoint significant G4 RNA signals, as corroborated by eCLIP experimental data.
We note that as the G4 confidence intensifies, there is an augmented inclination towards folded G4.
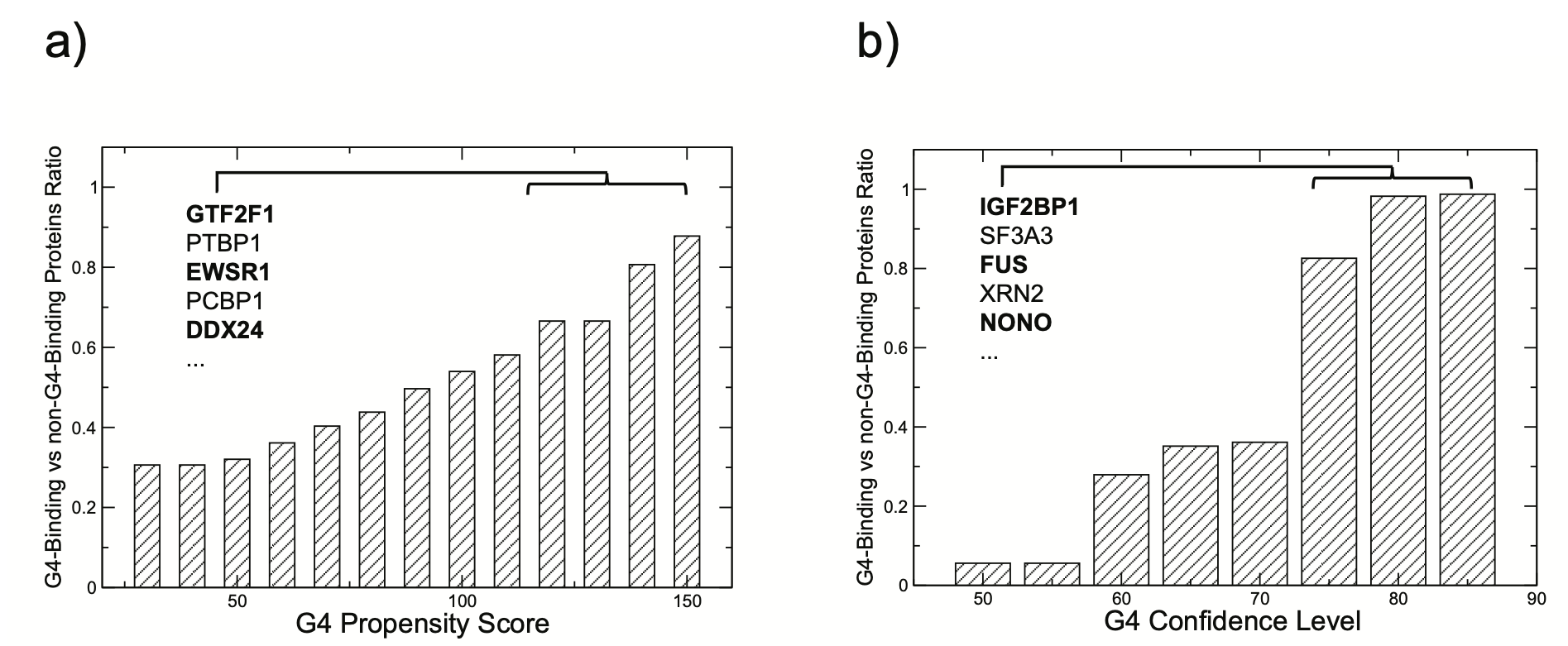
Legend a) Enhanced differentiation of G4 binders from non-binders is associated with rising G4 RNA propensity scores.
b) The differentiation intensifies with increased cleverMachine confidence levels. For both analyses, 'the G4-binding to non-G4-binding protein ratio is determined' by the count of G4s associated with each category.
Proteins predicted to bind folded G4 are written in bold.
Output interpretation
Please see the tutorial section of the documentation.